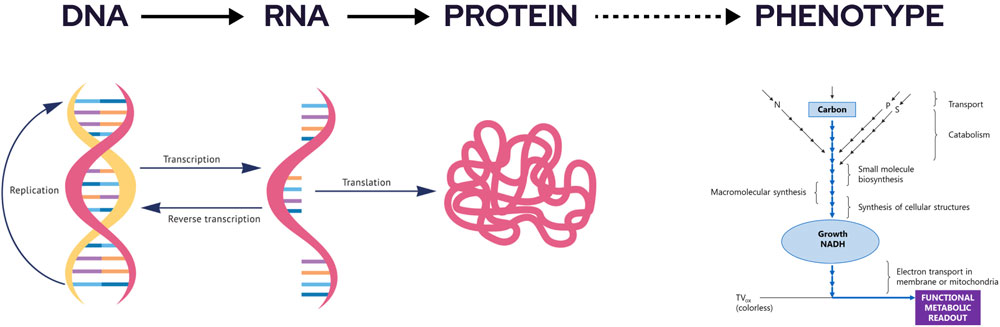
overview
DNA microarrays and proteomic methods made it possible to assay thousands of genes or proteins in a cell at once.
Phenotype MicroArray™ technology is the next step in characterization, making it possible to measure thousands of cellular phenotypes in a single experiment. Through comprehensive and precise quantitation of phenotypes, researchers can obtain an unbiased perspective of the effect on cells of genetic differences, environmental changes, and exposure to drugs or chemicals.
- Correlate functional phenotypes with genotypes
- Determine a cell strain’s metabolic and chemical sensitivity properties
- Elucidate drug susceptibility
- Optimize cell lines and bioprocess culture conditions
- Characterize cellular phenotypes for taxonomic or epidemiological studies
How it Works
Phenotype MicroArrays are preconfigured 96-well plates containing different classes of chemical compounds. After inoculation with a standardized cell suspension, they are designed to test for the functional presence or absence of thousands of phenotypes in a single experiment.
There are 10 panels designed to interrogate metabolic pathways along with ionic, osmotic and pH effects, and 10 panels to assess the sensitivity to various antimicrobials with different mechanisms of action.
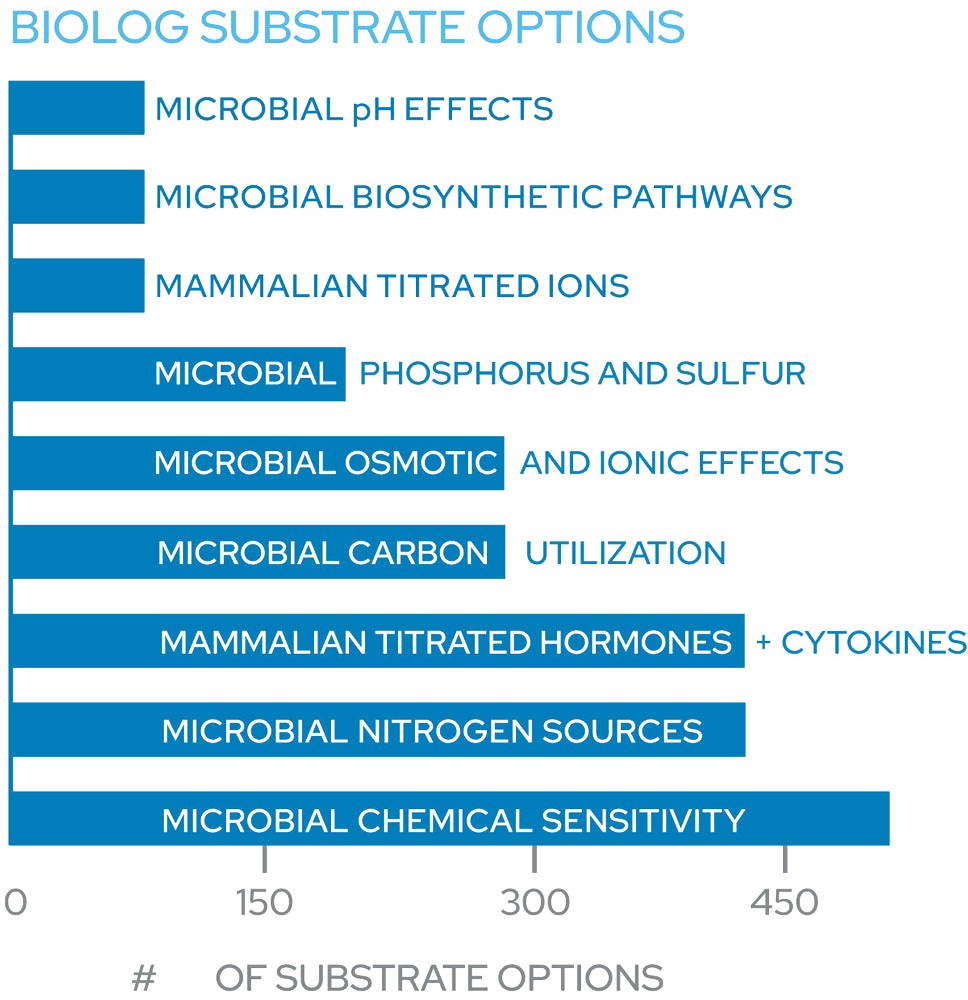
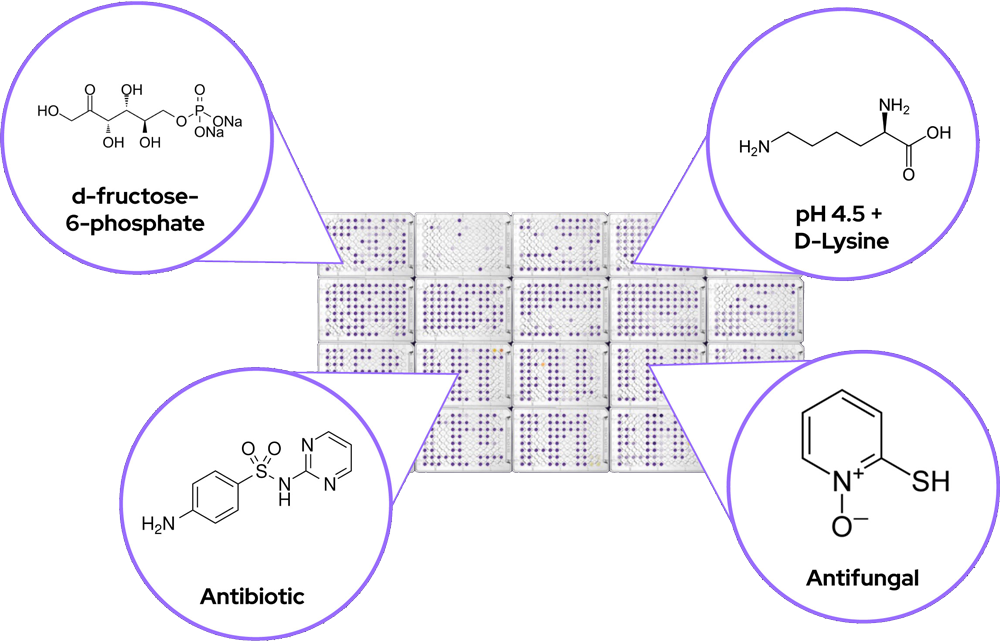

| Plate Name | Description |
|---|---|
| PM 1-2 Microplates | Carbon utilization assays |
| PM 3 Microplate | Nitrogen utilization assays |
| PM 4 Microplate | Phosphorus – Sulfur utilization assays |
| PM 5 Microplate | Biosynthetic pathway/nutrient stimulation |
| PM 6-8 Microplates | Nitrogen utilization assays |
| PM 9 Microplate | Osmotic/Ionic response assays |
| PM 10 Microplate | pH response assays |
| PM 11-20 Microplates | Bacterial chemical sensitivity assays |
| PM 21-25 Microplates | Yeast chemical sensitivity assays |

Technology
How Phenotype MicroArray Technology Works
Phenotype MicroArrays can be used in two ways. You can monitor cellular respiration by utilizing Biolog’s patented redox dye technology, which amplifies the signal from NADH production. You can also use the MicroArrays to monitor growth without the redox dye present.
Respiration and growth can be simultaneously monitored on Odin™, with and without the redox dye.
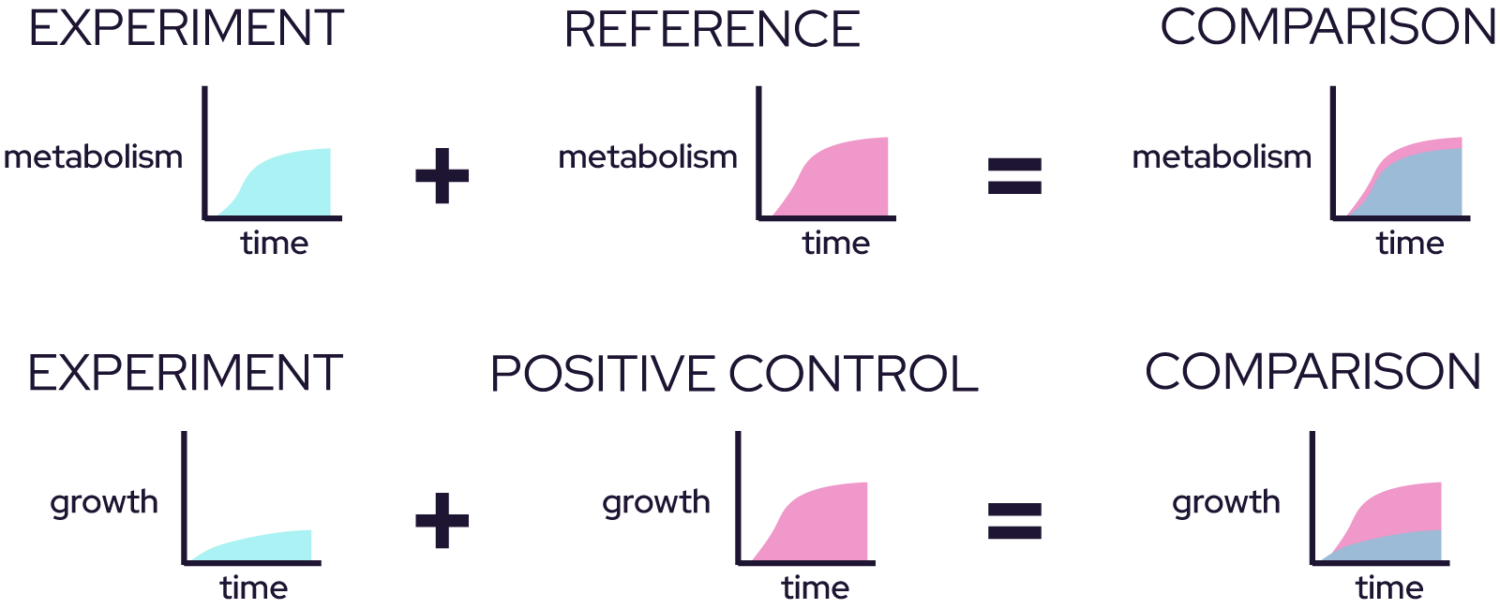
An example of how we compare phenotypes between two cell lines is depicted here. Cell line 1 (blue) and cell line 2 (yellow) are grown simultaneously under the same conditions. Our software will overlay the two respiration or growth curves to detect differences. The overlay area is green, allowing a qualitative view of the phenotypic differences between the cell lines.
Phenotypic Analysis with Odin
When using Phenotype MicroArrays, you need an instrument that will read at the right wavelengths and measure the changes kinetically over time at short intervals. You also need a software package to help you analyze all that data and identify the differences between your samples. With the Odin™ platform you can characterize phenotypes by monitoring growth curves and cell metabolism kinetics for microbial or mammalian cells for up to 50 plates at the same time. When combined with Phenotype MicroArrays, Odin can analyze up to 4,800 conditions unattended for hours or days.


Checking for Growth with odin
See what’s happening in two ways.
Odin measures a reporter dye over time at one wavelength to measure NADH production, effectively reporting the rate of metabolic respiration.
It also measures optical density (OD) at a second wavelength to determine how quickly the microbes are dividing. Taken together, you get a full picture of how your microbes grow best over time.
If you’re focused on cell growth, just leave the dye out. Odin automatically compares cell growth and metabolic activity curves to provide you with a clearer picture of how fast or slow your microbes are growing.

