Products
Biolog provides powerful cellular analysis tools for solving critical problems.



Products
Biolog provides powerful cellular analysis tools.
Biolog provides powerful cellular analysis tools for solving critical problems for pharmaceutical, food and beverage, industrial, and agricultural organizations and for publishable insights in academic research. Our patented technology is used in many applications, including microbial identification and characterization, as well as advanced phenotypic analysis for cells of all types, from microbial to mammalian.

Odin family
The all-in-one solution for phenotyping! Fully-automated metabolic characterization, growth kinetics, and microbial identification on one platform

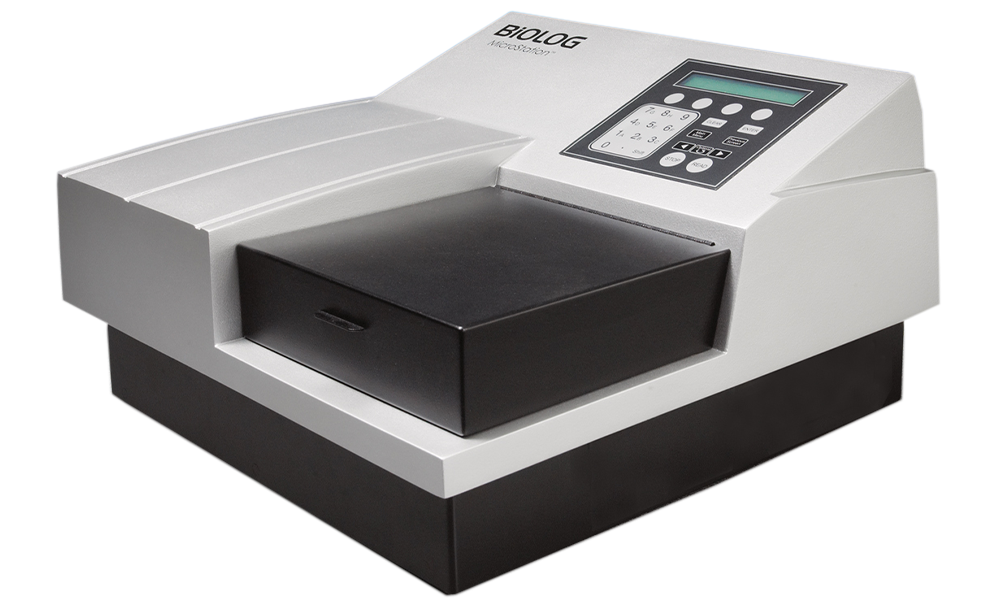
MicroStation
Semi-automated system for identification and classification of bacteria, yeast, and filamentous fungi

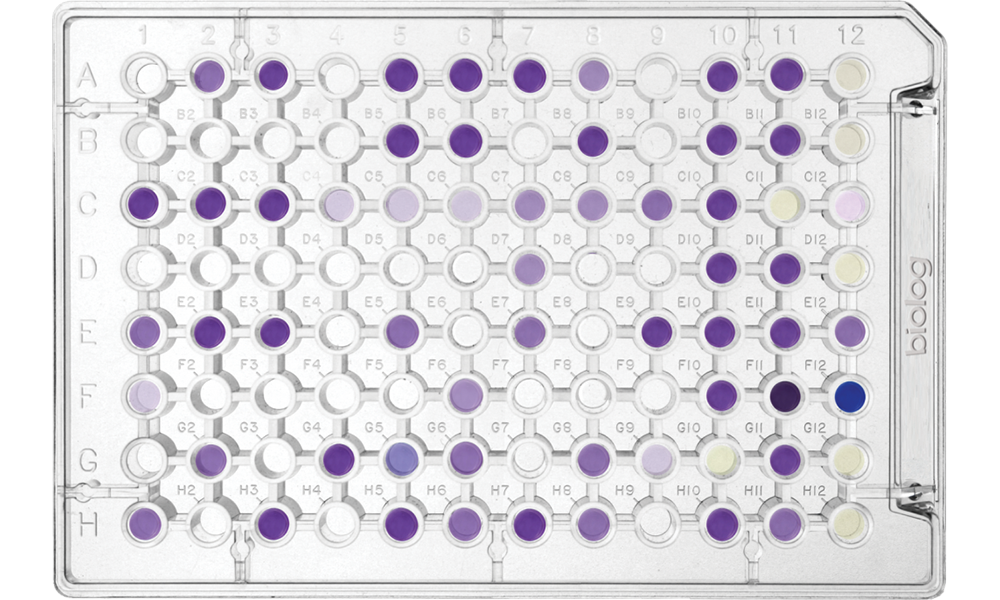
Phenotype MicroArrays
Microbial Phenotype MicroArrays
Screen nearly 2,000 specific cellular phenotypes in every experiment
Mammalian Phenotype MicroArrays
Investigate up to 1,400 metabolic and chemical sensitivity phenotypes of mammalian cells
Customizable Phenotype MicroArrays
Design your own phenotyping panel focused on your needs
MitoPlate Phenotype
Mitochondrial function assays with MitoPlates

Microbial Identification Microplates
Gen III Test Panel
Identify a broad range of gram-negative and gram-positive aerobic bacteria
Yeast Test Panel
Identify and characterize a broad range of yeast
Filamentous Fungi Test Panel
The first broad-based rapid identification and characterization product designed for filamentous fungi and yeast
Anaerobic Bacteria Test Panel
Identify a broad range of anaerobic and microaerophilic bacteria

Community Analysis Microplates
EcoPlate
Monitor environmental changes in microbial communities

Reagents
Microbial Detection using Chromogenic Media
Rainbow® Agars are a simple selective and chromogenic medium
Software Solutions
Odin Software
Acquisition and analysis data for Odin™ Platform for microbial characterization and identification.
Sherlock Software
Characterize your soil with Sherlock to automatically name all of the PLFA peaks.

